The emergence of antibiotic-resistant organisms is a major public health concern, particularly in hospitals and other health care settings.1,2 Antibiotic-resistant organisms appear to be biologically fit and are capable of causing serious, life-threatening infections that are difficult to manage because treatment options are limited. This increase in the prevalence of drug-resistant pathogens is occurring at a time when the discovery and development of new anti-infective agents is slowing down dramatically.2 Consequently, there is concern that in the not-too-distant future, we may be faced with a growing number of potentially untreatable infections.
In the past decade, several antibiotic-resistant pathogens have been identified as causes of serious infections among patients in hospital (Box 1). These organisms are typically resistant to multiple classes of antimicrobial agents and are therefore called multidrug-resistant organisms. Clostridium difficile, the major cause of diarrhea associated with antibiotic use, may also be considered a nosocomial antibiotic-resistant organism. In this article, we review important mechanisms of antibiotic resistance and discuss the clinical impact of key multidrug-resistant bacteria in hospital settings.
The genetics of antibiotic resistance
Genes can encode proteins or ribosomal RNA that enables bacteria to evade the actions of antibiotics. Such antibiotic resistance may either be intrinsic or acquired (Box 2).3 Intrinsic resistance is associated with the “usual” chromosomal genes or determinants of the organism's characteristics. Thus, an organism can be resistant to a specific class of antibiotics by virtue of its inherent genetic makeup. This form of resistance is predictable, which makes antibiotic selection straightforward. For example, all streptococci are intrinsically resistant to aminoglycosides (e.g., gentamicin and tobramycin), and all gram-negative bacilli are intrinsically resistant to vancomycin.
However, antibiotic resistance may also be acquired. This involves a change in the organism's genetic composition. This may occur by 1 of 2 mechanisms: there may be a mutation in the bacterial chromosomal DNA, or there may be acquisition of new genetic material. Mutations are generally uncommon events, perhaps occurring at a frequency of 1 event per 107–1010 bacteria, but may result in the development of resistance during therapy in organisms that are initially susceptible. An important example of this type of resistance is isoniazid resistance that can occur in Mycobacterium tuberculosis. This form of resistance is not transferable to other organisms. The probability of multiple resistance mutations occurring in a single organism is equal to the product of their individual probabilities. This is the rationale behind the use of combination therapy for the management of tuberculosis.
Perhaps of greater concern is the development of resistance because of the acquisition of new genetic material. Genes mediating antimicrobial resistance may be found on transferable segments of DNA such as plasmids, transposons or integrons (Box 2). Plasmids are extrachromosomal molecules of DNA that replicate independently from the bacterial chromosome. They may carry genes that convey resistance to antibiotics, as well as genes that may enhance bacterial fitness or virulence. Transposons carry antibiotic resistance genes along with genes that allow them to replicate and transpose, or “jump,” to other regions of the chromosome or to plasmids. An integron is a DNA structure capable of capturing genes. Although integrons are not themselves mobile, they may be carried by plasmids or transposons to other bacteria. These mobile genetic elements may be transferred from organism to organism, and even from one bacterial species to another. Multiple antibiotic resistance genes may be transferred at the same time. There are numerous examples of this type of resistance, including plasmid-mediated production of β-lactamase enzymes, which are capable of inactivating penicillins or cephalosporins in Staphylococcus aureus, Escherichia coli or Enterobacter species.
Mechanisms of antibiotic resistance
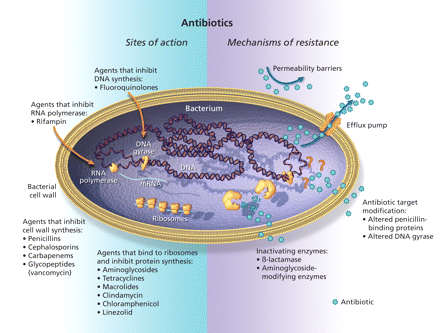
Antibiotics work by interacting with specific bacterial targets, inhibiting bacterial cell-wall synthesis, protein synthesis or nucleic acid replication. To accomplish this, the antibiotic must have access to and bind to its bacterial target site (Figure 1). Whether antibiotic resistance is intrinsic or acquired, the genetic determinants of resistance encode specific biochemical resistance mechanisms that may include enzymatic inactivation of the drug, alterations to the structure of the antibiotic target site, and changes that prevent access of an adequate concentration of the antimicrobial agent to the active site (Table 1).3
Figure 1: Sites of action and potential mechanisms of bacterial resistance to antimicrobial agents. Modified with permission from the American Association for the Advancement of Science (>Science 1992;257:1064–73).3 Image by: Lianne Friesen and Nicholas Woolridge
Table 1.
Enzymatic drug inactivation
Bacteria may produce enzymes that modify or destroy the chemical structure of an antibiotic, which renders it inactive. This mechanism of resistance is probably best exemplified by the β-lactamase family of enzymes, which act by hydrolyzing the β-lactam ring of penicillins, cephalosporins and carbapenems. There are hundreds of β-lactamase enzymes that may be distinguished by their substrate profiles and activities. Some β-lactamase genes are chromosomal, whereas others are located on plasmids or transposons. Penicillin resistance in S. aureus and Neissseria gonorrheae, ampicillin-resistance in Haemophilus influenzae, and resistance to extended-spectrum cephalosporins in E. coli and in Enterobacter species are all commonly mediated by the production of β-lactamases.
Resistance to extended-spectrum cephalosporins (e.g., cefotaxime, ceftriaxone, and ceftazidime) has arisen primarily by 1 of 2 mechanisms, both of which involve the production of β-lactamases.4 In E. coli and Klebsiella species, plasmid-mediated extended-spectrum β-lactamases have most often emerged as mutations of the TEM, SHV or CTX-M genes (Ambler class A β-lactamases). In the past decade, the CTX-M type of extended-spectrum β-lactamases has become predominant in many parts of the world, including Canada.5,6 In other gram-negative bacilli, such as Enterobacter, Citrobacter or Serratia species, this resistance most often arises because of selection of mutants that overproduce chromosomally encoded AmpC cephalosporinases (Ambler class C β-lactamases), although plasmid-mediated ampC genes have also been identified.7 Both resistance mediated by Ambler class A extended-spectrum β-lactamases and AmpC are associated with cross-resistance to penicillins and cephalosporins. In addition, these bacteria are often also resistant to other antibiotic classes, such as fluoroquinolones, trimethoprim-sulfamethoxazole and aminoglycosides.
Broad-spectrum β-lactamases, called carbapenemases, are capable of degrading carbapenem antibiotics and may be responsible for resistance to imipenem and meropenem in Pseudomonas aeruginosa and other gram-negative bacilli. An important and rapidly emerging group of carbapenemases is the KPC (Klebsiella pneumoniae carbapenemase) family of enzymes, which are capable of inactivating all β-lactam drugs, including carbapenems.8 The KPC genes are found on plasmids that often carry determinants of resistance to other classes of antimicrobials. Although these enzymes are most commonly found in K. pneumoniae, they have spread to other Enterobacteriaceae. To date in Canada, there have been no reports of infections caused by organisms with K. pneumoniae carbapenemase enzymes, but outbreaks caused by these multidrug-resistant strains have been reported in the United States and in many countries in Europe, Asia and South America.8,9
Alteration of the antibiotic target site
For antibiotics to work, they must bind to a specific bacterial target site, which varies depending on the class of antibiotic. A change in the structure of the target may result in the inability of the antibiotic to bind to its target. For example, β-lactam antibiotics act by binding to structures in the bacterial cell wall called penicillin-binding proteins. Methicillin-resistant strains of S. aureus (MRSA) possess a genetic element called staphylococcal cassette chromosome mec (SCCmec), which contains the mecA gene that codes for the production of an altered penicillin -binding protein (PBP2a) that does not effectively bind β-lactam antibiotics.10 As a result, MRSA is resistant to all of the currently available penicillins, cephalosporins and carbapenems. Changes in penicillin-binding proteins also account for penicillin resistance in Streptococcus pneumoniae.11
Another example of antimicrobial resistance caused by an altered target site is resistance to fluoroquinolones (e.g., ciprofloxacin, levofloxacin and moxifloxacin). Fluoroquinolones act by inhibiting proteins called DNA gyrases (encoded by gyrA and gyrB genes) and topoisomerases (encoded by parC and parE), which are essential for bacterial DNA replication. Mutations in specific regions of the gyrA or parC genes (known as the quinolone-resistance determinant region) result in alterations to DNA gyrase or topoisomerase, and therefore result in an altered target binding site.12 Typically, multiple step-wise mutations must occur for resistance to fluoroquinolones to emerge.
Prevention of antibiotic access to the target site
For an antibiotic to bind to its target, it must arrive at its target site and achieve an adequate concentration. Therefore, another strategy that has evolved to counteract antimicrobial activity is to prevent access of the drug to its target site. This may occur because of a permeability barrier or because of the presence of an efflux pump mechanism.
The cell wall of gram-negative bacteria consists of inner and outer membranes that act as a permeability barrier. To allow movement of essential compounds through the outer membrane, the bacterial cell produces outer membrane proteins (porins), which allow diffusion of molecules, including antibiotics into the cytoplasm. Mutations that cause changes in the structure of the outer membrane proteins can result in a permeability barrier that impedes access of antimicrobial agents to their active site. This mechanism may account for resistance to β-lactams and aminoglycosides in P. aeruginosa and other gram-negative bacilli.13
Instead of preventing penetration of antibiotics to the active site, some organisms have evolved an active efflux mechanism that pumps out antibiotics from the cytoplasm before they can bind to their target.14 These efflux pumps have been found in both gram-positive and gram-negative organisms. Such pumps may be responsible for resistance to tetracyclines, macrolides (e.g., erythromycin), clindamycin, fluoroquinolones and aminoglycosides. Some pumps may be specific for only 1 class of antibiotics, but others may be associated with resistance to multiple drugs because they are capable of exporting many different classes of antimicrobials.
The emergence and transmission of antimicrobial-resistant organisms
A drug-resistant organism may be introduced into a health care facility with the admission of a patient who is infected with or who has been colonized by such a strain. Alternatively, antimicrobial resistance may emerge in bacteria as a response to selective antibiotic pressure, or a resistant organism may spread from person to person (Table 2). Often, a combination of these factors may be involved in the emergence and transmission of antimicrobial resistance within a health care facility.
Table 2.
Selective antibiotic pressure
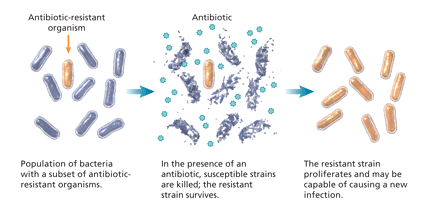
Selective pressure refers to the environmental conditions that allow organisms with certain characteristics to survive and proliferate. Exposure to an antibiotic, for example, may inhibit or kill the majority of the bacterial population who are susceptible. However, a resistant subset of organisms may not be inhibited or killed by the antibiotic (Figure 2). These bacteria may be intrinsically resistant to the antibiotic, or they may have acquired resistance. Thus, antimicrobial use selects for the emergence of resistant strains of organisms that may then proliferate and become predominant.15 Indeed, antimicrobial resistance in health care facilities and the community is largely determined and magnified by the selective pressure of antimicrobial use.16
Figure 2: Effect of selective antibiotic pressure in bacteria. Image by: Lianne Friesen and Nicholas Woolridge
There are numerous examples that illustrate the direct relation between antimicrobial use, both appropriate and inappropriate, and antimicrobial resistance at both the population and individual patient level. In Finland, a significant association was found between macrolide consumption and macrolide resistance rates for group A Streptococcus.17 Similar correlations have been identified for macrolide use and resistance in S. pneumoniae isolates in the United States,18 and for fluoroquinolone use and decreased susceptibility of pneumococci to fluoroquinolones in Canada.19 In intensive care units and other hospital settings, increased use of fluoroquinolones is correlated with a greater incidence of ciprofloxacin resistance in gram-negative bacilli.20–22 Similarly, many studies have documented prior antimicrobial exposure as a significant risk factor for subsequent acquisition of an antibiotic-resistant organism at the patient level. For example, prior exposure to broad-spectrum cephalosporins has been associated with acquisition of vancomycin-resistant enterococci and organisms that produce extended-spectrum β-lactamases.23,24 Previous fluoroquinolone or carbapenem use is an important risk factor for infection due to gram-negative organisms resistant to fluoroquinolones or carbapenems.25–27
However, the relation between antimicrobial use and resistance is complex. In a study involving 8 hospitals in the United States, high intensity of antimicrobial use was not necessarily associated with high rates of resistance.28 Moreover, facility-level and individual risk factors for antibiotic resistance may be different, especially with regards to antimicrobial exposure. In a parallel analysis of individual and aggregated data on antibiotic exposure and resistance, different results were obtained with facility-level and individual patient-level analyses.29 In an ecologic facility-wide analysis, there was no apparent relation between intensity of antibiotic use and rates of resistance. But when the same data were analyzed at the individual patient level, there were significant associations between antibiotic exposure and resistance.
Clonal transmission of antibiotic-resistant organisms
Organisms resistant to antimicrobial agents can be spread from patient to patient in health care facilities, often via the contaminated hands of health care personnel, contaminated medical or surgical equipment, or the inanimate hospital environment. This type of spread is generally clonal, involving the transmission of a single strain of the antibiotic-resistant organism. Outbreaks caused by clonal transmission of an antimicrobial-resistant organism have commonly been reported for MRSA, vancomycin-resistant enterococci, C. difficile and multidrug-resistant gram-negative bacilli.24,30–35 Clonal outbreaks at multiple institutions may also occur with transmission of a common strain in multiple health care facilities, even in diverse geographic regions.30,33,34,36 However, clonality is less likely to occur with sporadic disease in areas with a relatively low prevalence, or with certain organisms and mechanisms of resistance.37–39 In some settings, an outbreak of antimicrobial resistance may occur from transmission of a plasmid rather than dissemination of a single resistant strain.40
Selected antibiotic-resistant organisms in hospitals
Methicillin-resistant Staphylococcus aureus
Although there is considerable variation in the rates of MRSA from country to country, and even from hospital to hospital within a country, MRSA is currently the most commonly identified antibiotic-resistant pathogen among patients in hospital.41 Rates of MRSA infections have increased in both US and Canadian hospitals; however, the rates are much higher in the United States.42,43 Until recently, MRSA was considered to be primarily a nosocomial pathogen, affecting older adults with comorbidities in hospital or long-term care settings. However, in the past decade, community-associated MRSA (CA-MRSA), which involves a small number of unique MRSA strains (clones), has emerged in many parts of the world, including Canada.44–46 Patients affected by CA-MRSA often do not have commonly recognized health care–associated risk factors. CA-MRSA may cause infections at any site but are most often associated with skin and soft tissue infections, including pustulosis, furunculosis and abscesses. Invasive disease and fatal necrotizing pneumonia have also been reported.47 Not surprisingly, CA-MRSA strains have been introduced into health care settings and have been associated with hospital-acquired bacteremia, surgical-site infections and outbreaks in maternity units and hospital nurseries.48
Vancomycin-resistant Enterococcus
Although generally less virulent than S. aureus, Enterococcus species (e.g., Enterococcus faecalis, Enterococcus faecium) may also cause serious, life-threatening infections. Resistance to vancomycin in enterococci is caused by synthesis of modified cell-wall precursors that do not bind glycopeptides. This occurs with the acquisition of a plasmid-associated gene cluster, most commonly the vanA or vanB genes.49 These genes are transferable and may spread from enterococci to MRSA, thereby further complicating the treatment of infections caused by this organism.50 In Canada, 15% of inpatients with vancomycin-resistant enterococci were found to also be colonized with MRSA.51 Prior antimicrobial therapy with oral vancomycin, broad-spectrum cephalosporins or metronidazole has been identified as an important risk factor for vancomycin-resistant enterococcal infection or colonization.23,52 Environmental contamination with vancomycin-resistant enterococci or exposure to contaminated medical equipment may also contribute to nosocomial transmission.53
Multidrug-resistant gram-negative bacilli
Multidrug-resistance in gram-negative bacilli is generally defined as resistance to more than 2 classes of antimicrobial agents. Typically, multidrug-resistant gram-negative bacteria are resistant to penicillins (including those combined with a β-lactamase inhibitor), cephalosporins, fluoroquinolones, trimethoprim-sulfamethoxazole and aminoglycosides. However, some strains may also be resistant to the carbapenems, often leaving colistin as the only agent available for treatment of these infections.54
The major multidrug-resistant gram-negative pathogen in most Canadian hospitals is P. aeruginosa, which is most often isolated from patients in intensive care units. At present, about 30% of P. aeruginosa isolates from intensive care units in the United States and Canada are resistant to fluoroquinolones.42,55,56 In Canadian intensive care units, about 13% of isolates were found to be resistant to multiple antibiotics.56 Carbapenem-resistance was identified in 14% of P. aeruginosa isolates from intensive care units in Canada.56 About 21% of P. aeruginosa isolates from intensive care units in the United States were resistant to carbapenems.42
Acinetobacter baumannii is also an important nosocomial pathogen acquired in intensive care units. This pathogen may cause pneumonia or urinary tract, wound, or bloodstream infections. Acinetobacter are generally resistant to most classes of antimicrobials, leaving carbapenems, and possibly glycylcyclines (tetracycline derivatives such as tigecycline), as the only effective drugs. The emergence of carbapenem-resistant A. baumannii in many parts of the world is disturbing and poses a threat to the effective management of these infections.57 Multidrug-resistant A. baumannii is identified infrequently in Canadian hospitals, although an outbreak was reported in a burn unit of a tertiary care hospital in Toronto.58 Nosocomial outbreaks due to multidrug-resistant A. baumannii that originated with injured military personnel returning from Afghanistan and Iraq have recently been reported in the United States and the United Kingdom.59 Similarly, a number of injured Canadian soldiers returning from active duty in Afghanistan have received treatment for multidrug-resistant Acinetobacter respiratory and wound infections in Canadian hospitals.60
Clostridium difficile
C. difficile is the major cause of diarrhea associated with antibiotic use and is the most common infectious cause of nosocomial diarrhea. A major outbreak of C. difficile infection occurred in many hospitals in Quebec beginning in 2002.32,61 Markedly increased disease rates (as high as 156 cases per 100 000 people) and severity occurred, especially among elderly people. In this outbreak, there was often a poor response to metronidazole therapy.61,62 The emergence of such severe disease is thought to have occurred because of the presence of a hypervirulent epidemic strain of C. difficile, known as PCR ribotype 27, or North America pulso-type 1 (NAP1).63 The same strain of C. difficile has caused extensive and severe disease in the United States and Europe.33 It is not clear why this strain appeared and caused such severe disease in the past few years, but this may in part be related to changing patterns of antimicrobial use in hospitals. The NAP1 strain associated with these outbreaks is resistant to fluoroquinolones, and fluoroquinolone use was found to be a major risk factor for C. difficile-associated disease in the Quebec outbreak.32 The NAP1 strain has now been identified in most provinces, and it has become the predominant strain in many hospitals, indicating the potential for severe outbreaks in many parts of the country.
Impact of antimicrobial resistance
The burden of antimicrobial resistance refers to the impact of events that would not have occurred if resistance had not been present. These outcome measures may include excess mortality, morbidity (e.g., length of hospital stay and complications) and attributable costs (e.g., costs to the hospital, patient and society). The risk of such adverse outcomes has been found to be higher in patients with infections caused by an antibiotic-resistant organism compared with infections caused by susceptible strains of the same pathogen, even after adjustment for underlying comorbidities.64–68 For example, in a meta-analysis performed to examine the impact of methicillin-resistance on mortality among patients with S. aureus bloodstream infections, a significant increase in mortality (nearly a double increase in risk) was found to be associated with MRSA bacteremia when adjusted in a random-effects model (odds ratio 1.93, 95% confidence interval 1.54–2.42, p < 0.001).69 In a multivariable model, MRSA bacteremia was associated with increased length of hospital stay (1.29-fold increase, p = 0.016), with an attributable length of stay of 2 days.64 Similarly, attributable prolonged excess hospital stay was identified for infections caused by resistant strains of extended-spectrum ß-lactamase-producing K. pneumoniae (median post-infection hospital stay of 11 days for susceptible strains compared with 29 days for resistant strains, p = 0.03) and for carbapenem-resistant P. aeruginosa (20 days v. 34 days, p = 0.002).68 Several studies have attributed these adverse outcomes to delays in instituting effective antimicrobial therapy and use of inadequate initial therapy.70,71
Not surprisingly, adverse outcomes and prolonged hospital stays associated with antimicrobial-resistant organisms have been associated with increased costs. However the economic burden of antimicrobial resistance on the health care system has not been well defined. Thirteen years ago in the United States, cost estimates for managing antimicrobial resistance were between $0.1 billion and $10 billion per year.72 In 2001, the mean attributable cost associated per patient with MRSA infection in a Canadian hospital was $14 360.73 The incremental cost for care of patients with enterococci in Canadian hospitals was estimated to be about $6700 per patient.74 Incremental costs were attributed to length of hospital or intensive care unit stay, lost revenue from private isolation rooms, need for more expensive antibiotics and increased laboratory testing, and the added expense of infection control interventions, including the use of antiseptic soaps, gloves and gowns.
Summary
The burden of antibiotic resistance continues to increase and is acknowledged to be a major threat to the treatment of infectious diseases, particularly among patients in hospital. The reasons for the variability in resistance rates around the world and within different parts of Canada is unknown but may be related to differences in diagnostic procedures, patterns of antimicrobial use or infection prevention and control practices.
Other important gaps in our knowledge include uncertainty about how understanding specific mechanisms of resistance may lead to the identification of novel targets for new antimicrobial drug development. A better understanding of the relative importance of selective pressure related to antibiotic use compared to cross-infection as mechanisms for emergence and spread of antimicrobial resistance would also be important to design and evaluate effective infection prevention and control strategies.
Infections caused by antimicrobial-resistant organisms are almost always associated with increased attributable mortality, prolonged hospital stays and excess costs. Although the rates of antibiotic resistance in Canada tend to be lower than those in many other parts of the world, complacency would be a mistake. Rates of antibiotic-resistant organisms have increased in Canadian health care facilities in the past 10 years, and the rates will continue to increase unless aggressive control measures are implemented.75 These interventions must include enhanced surveillance of antibiotic resistance, attention to hand hygiene and other standard infection prevention and control measures, and antibiotic stewardship to ensure appropriate use of antimicrobial agents.76,77
@@ See related commentary by Nicolle and colleagues, page 371, and related analysis paper by Patrick and Hutchinson, page 416
-
The incidence and adverse consequences of infections caused by antimicrobial-resistant organisms continue to increase.
-
Acquired antibiotic resistance may arise from bacterial chromosomal DNA mutations or as a result of transfer and acquisition of new genetic material.
-
The major mechanisms of antimicrobial resistance include enzymatic inactivation of the drug and bacterial changes that prevent access or binding to the drug's target.
-
The emergence and spread of antimicrobial-resistant organisms within health care facilities or the community may be related to clonal transmission of a common “epidemic” strain.
-
Emergence and spread may be promoted by antibiotic selection pressure.
-
Enhanced infection prevention and control strategies need to be developed, implemented and evaluated to stem the rise of infections caused by antimicrobial-resistant organisms.
Key points
Footnotes
-
This article has been peer reviewed.
Contributors: Both of the authors contributed to the conception of this review, and participated in reviewing and analyzing the literature. Both of the authors drafted portions of the review, revised it critically and approved the version submitted for publication.
Competing interests: None declared.
REFERENCES
- 1.↵
- 2.↵
- 3.↵
- 4.↵
- 5.↵
- 6.↵
- 7.↵
- 8.↵
- 9.↵
- 10.↵
- 11.↵
- 12.↵
- 13.↵
- 14.↵
- 15.↵
- 16.↵
- 17.↵
- 18.↵
- 19.↵
- 20.↵
- 21.
- 22.↵
- 23.↵
- 24.↵
- 25.↵
- 26.
- 27.↵
- 28.↵
- 29.↵
- 30.↵
- 31.
- 32.↵
- 33.↵
- 34.↵
- 35.↵
- 36.↵
- 37.↵
- 38.
- 39.↵
- 40.↵
- 41.↵
- 42.↵
- 43.↵
- 44.↵
- 45.
- 46.↵
- 47.↵
- 48.↵
- 49.↵
- 50.↵
- 51.↵
- 52.↵
- 53.↵
- 54.↵
- 55.↵
- 56.↵
- 57.↵
- 58.↵
- 59.↵
- 60.↵
- 61.↵
- 62.↵
- 63.↵
- 64.↵
- 65.
- 66.
- 67.
- 68.↵
- 69.↵
- 70.↵
- 71.↵
- 72.↵
- 73.↵
- 74.↵
- 75.↵
- 76.↵
- 77.↵