New mechanisms of antibiotic resistance
In the 1970s the bacterium Acinetobacter baumannii was sensitive to most antibiotics; but over the last 35 years, some A. baumannii strains have become resistant to virtually all antibacterial drugs. In some countries, infections with this organism have become a public health problem: A. baumannii is responsible for up to 10% of all gram-negative infections in intensive care units in Europe.
So what has changed?
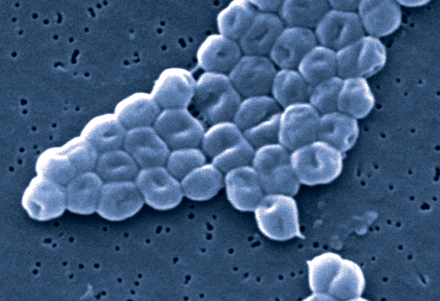
Figure. A scanning electron micrograph of a cluster of gram-negative, nonmotile Acinetobacter baumannii bacteria, highly magnified (× 165 000). Photo by: Janice Carr
Pierre-Edouard Fournier and colleagues recently attempted to answer this question when they compared the genomes of 2 strains of A. baumannii: one from an outbreak in France that is resistant to virtually all antibiotics versus one associated with human body lice that is extremely susceptible (PLoS Genetics 2006;2(1):e7). The resistant strain had caused 26% of the people infected with it to die (J Clin Microbiol 2003;41:3542-7).
The scientists discovered that the genome of the resistant strain of A. baumannii possesses a chromosomal region — a so-called resistance island — containing 45 genes that contribute to antibiotic resistance. Many of these genes are of types not seen before, and their various roles in antibiotic resistance still need to be verified.
Analysis of these “resistance” genes revealed similarities with species of Pseudomonas, Escherichia and Salmonella, which suggests that exchanges of genetic information have likely occurred. Since all of these bacteria are commonly found in aqueous environments in hospitals, Fournier and colleagues suggest that antimicrobial pressure in such settings probably promotes genetic exchanges between and among these pathogens. Moreover, the areas around the resistance genes seem to be primed to capture additional genes, which perhaps explains why the organism has been so quick to develop drug resistance.
Antibiotic resistance in human pathogens is a major clinical problem; genome comparisons will illuminate some aspects of how this occurs. Our understanding of these mechanisms is crucial to the development of novel agents to overcome these pathogens' defenses. — Compiled by David Secko, Vancouver








