Personalized care is the careful tailoring of investigations and treatments to individual patients based on evidence, consideration of circumstances and clinical skill.
Precision medicine relies on increasingly detailed molecular characterization of disease states using the biologic omics platforms to better individualize diagnostics, prognostics and therapeutics.
Although there have been impressive advances in precision medicine in the past decade (particularly in the field of oncology), the potential effects of precision medicine may be overestimated.
Better health outcomes require advances not only in omics-based medicine but also in traditional elements of personalized care.
The term precision medicine refers to the tailoring of diagnostics or therapeutics to individual patients based on their unique genetic and physiologic characteristics.1 Although “personalized or individualized care” (i.e., tailoring investigations and therapies to each patient) has long been the hallmark of good clinical care, more recently, an emphasis on genomics, proteomics and other biologic omics platforms has come to be understood to underpin personalized care. Some commentators conflate precision medicine and personalized care, but they are not synonymous.
One key criticism of evidence-based medicine is that even efficacious therapies benefit only a minority of patients to whom they are administered, but all treated patients are exposed to the costs and potential harms of those therapies.2–4 In 2017, we still conduct randomized controlled trials (RCTs) in thousands of people of similar phenotype to detect small to moderate treatment effects and then extrapolate the average response seen in the trials to patients who are often older, sicker, have more comorbidities and take more medications than trial participants. As a result, the potential effect of any intervention is often uncertain in a specific patient. Precision medicine has been advanced as a potential solution to this problem.
Why is personalized care valuable?
An example of current personalized care by clinicians is the use of biomarkers (such as brain natriuretic peptide for patients with heart failure or serum troponin for those with chest pain) to better delineate disease subtype, define prognosis or to inform therapy decisions. Clinicians also practice personalized care by reviewing eligibililty criteria for trials and tables of baseline characteristics to extrapolate clinical evidence when treating a specific patient either by gestalt or by applying the findings of subgroup analyses within trials or systematic reviews, for example, or using clinical prediction rules and formal decision analyses.5 These efforts are imperfect because subpopulations are often defined by only one or two characteristics, and any patient in clinical practice may fit into several subgroups with discordant treatment effects.
Precision medicine holds promise for better personalization of care in the future; however, an increasing focus on molecular diagnostics and imaging may divert attention away from the traditional cornerstone of clinical medicine — personomics. Personomics involves determining “an individual’s unique life circumstances that influence disease susceptibility, phenotype, and response to treatment,”6 which includes consideration of social determinants of health as well as clinical features. For example, a clinician may exercise more caution when increasing the dose of antihypertensive drugs for a patient who is frail, at risk of falls and who lives alone than for an otherwise healthy person. In addition, numerous nondisease factors influence a clinician’s prescription choices, including a patient’s adherence patterns, lifestyle behaviours, ability to afford specific drugs and whether or not they can comply with frequent laboratory testing.
Clinicians quickly learn that making diagnoses and defining treatment options are only the first steps in caring for a patient. Developing rapport between patient and clinician is vital not only for identifying the severity of symptoms but also for placing the disease/symptom complex in the context of that patient’s values, resiliency and social circumstances. A recent narrative review of approaches to the management of common symptoms reported that up to one-third of common symptoms seen in primary care do not have a clear-cut disease explanation.7 In the context of clinical uncertainty, trust between a patient and an engaged clinician can be therapeutic. Continuity and rapport with a clinician is an important influence on patient adherence to physicians’ recommendations8 — the importance of this should not be underestimated, because adherence is a major driver of efficacy, safety and cost-effectiveness of treatments. The clinical evaluation also plays a key role in the diagnostic cascade: a large prospective study from one hospital estimated that more than three-quarters of all diagnoses ever made are established by the initial history and physical exam.9 Similar findings from primary care settings10 and specialist clinics11 highlight the importance of clinical evaluation in the diagnosis and treatment of illness. Furthermore, both subsequent investigations and the interpretation of many diagnostic tests (e.g., exercise stress tests or pulmonary scintigraphy) depend on the clinical context.
What can precision medicine offer?
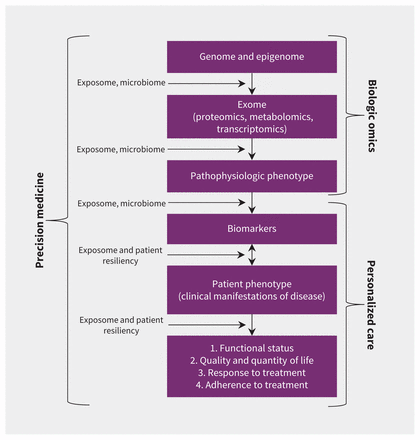
Precision medicine represents an extension of traditional personalized care through more precise individualization of diagnosis, prognosis and therapy estimates for each patient by using sophisticated molecular diagnostics and imaging made possible by recent technologic advances (Figure 1).
Although simplified for presentation, each box in the figure is interconnected with other layers in a bi-directional fashion. In addition to the connections between layers, external influences (denoted in the column on the left-hand side) can affect layers and the influence of one layer on others. The exposome encompasses all exogenous and endogenous exposures arising from the physical environment, patient lifestyle behaviours and the other social determinants of health. Patient resiliency encompasses the host of psychosocial factors that influence patient symptoms and their individual response to disease.
Some advances in diagnostics are impressive and encouraging. For example, a number of uncommon diseases (e.g., cystic fibrosis and sickle cell anemia) have now had their molecular diagnosis established by gene sequencing — enabling screening for late-onset fatal disease (such as Huntington disease), prenatal counselling and the hope that eventually curative gene-targeted therapies may be developed. As of June 30, 2017, the National Institutes of Health Genetic Testing Registry listed 51 519 genomic tests for 10 708 uncommon conditions (available at www.ncbi.nlm.nih.gov/gtr/). In addition, genomics has been pivotal in identifying the source and specificity of pathogen strains responsible for outbreaks (e.g., Clostridium difficile or tuberculosis).12,13 In the near future, genomic sequencing of pathogens may permit rapid characterization and earlier institution of specific antibiotic therapies in sepsis (thus obviating the current need for broad-spectrum antibiotics for 48 hours while waiting for results for culture and sensitivity tests).2
Other advances in genomics allow for the identification of individuals at risk for certain diseases, such as breast and ovarian cancers associated with BRCA1 and BRCA2 genes, and 16 other cancers in the National Institutes of Health Cancer Genome Atlas (available at www.cancergenome.nih.gov). Ongoing large cohort studies that collect detailed phenotypic data and outcomes, and include whole genome sequencing of participants, as well as characterization of their microbiomes and proteomic profiles, may result in expansion of the number of diseases for which screening and early treatment is feasible.2 As an extension of this concept, many RCTs (e.g., the ongoing VICTORIA [Vericiguat in Participants with Heart Failure with Reduced Ejection Fraction] Trial, NCT02861534) now include collection of novel biomarkers and biosamples for omic diagnostics at baseline to explore pathophysiologic reasons for any heterogeneity in treatment responses and, ultimately, to better define treatment-responsive subgroups.
Genotype-guided prescribing is also increasingly common. Because therapies shown to be efficacious in RCTs benefit only a few patients to whom they are administered but expose all to cost and potential harm, better targeting of interventions to those most likely to benefit and least likely to be harmed is a laudable goal. Careful genomic research has resulted in changes to recommendations for treatment regimens in some guidelines — for example, for patients with colon cancer who do or do not have KRAS gene mutations,14 and for patients with breast cancer with different HER2 status.15 The US Food and Drug Administration has approved almost two dozen new drugs that target specific oncogenes in the past five years.2 It is possible that pharmacogenomics will have an even larger effect with respect to drug dosing: 18% of outpatient prescriptions in the United States are for drugs linked to genetic variants that substantially influence their clinical effect (with upwards of three- to fourfold differences in treatment effectiveness), including antidepressants, antivirals, statins, antiepileptics, some antiplatelet agents and analgesics.2,3
Striking a balance
Although pharmacogenomics holds substantial promise, we are still in the early days of bespoke therapies in medicine: current estimates suggest that fewer than two in 100 patients with cancer may benefit from efforts in precision oncology.16 Although the Clinical Pharmacogenetics Implementation Consortium (available at www.cpicpgx.org) identifies over 350 gene–drug interactions, only a few have been endorsed for genotype-guided prescribing by the Centers for Disease Control and Prevention Office of Public Health Genomics owing to a paucity of outcome studies.17–19 Results from several randomized comparisons of pharmacogenomic-directed drug therapy against usual care have been disappointing.19 For example, a meta-analysis of nine trials showed no statistically significant benefits (for time in therapeutic range, bleeding or thromboembolic events) with genotype-guided dosing of warfarin.20
Evidence addressing the effect on safety and efficacy of precision medicine is scarce, and few studies have explored its value proposition. A systematic review published in 2014 found only 59 cost–utility analyses of current precision medicine tests — 20% were cost saving (largely by identifying patients in whom costly treatments should be avoided).21
Decoding each patient’s genome is unlikely to help much with the personalization of care for several reasons. First, genetic risk variants have small effect sizes (odds ratios of 1.5 or less), and most common diseases (e.g., diabetes and hypertension) have multifactorial causes.22 In a review of comparative effectiveness studies, Phillips and colleagues concluded that testing for genetic variants adds little to traditional algorithms for risk prediction, 19 and behavioural factors (e.g., smoking, activity levels and obesity) overwhelm any excess risk attributable to genetics because socioeconomic and lifestyle factors are still the main drivers of premature death in the developed world.23 Second, some genetic risk variants have identified increased risk for diseases for which no preventive therapies are currently available (e.g., the apoɛ4 allele has been associated with an increased risk of Alzheimer disease24). Third, many genomic variations appear to have no or minimal influence on health but may prompt a costly cascade of investigations and potential treatments in otherwise healthy people. For example, an exploratory study involving 12 adult participants reported that whole genome sequencing in healthy individuals identified a median of five personal disease-risk findings, which prompted a panel of clinicians to recommend three or more expensive diagnostic tests or referrals per healthy individual.25 Therefore, whole genome sequencing in individual patients could lead to increased costs for the public purse, even if patients paid for it privately through initiatives such as 23 and Me.26 Fourth, contrary to what might be expected, patients do not always modify their behaviour even when they are informed that they are in a genetic high-risk group. Instead, they often have increased anxiety and generate more costs through more frequent encounters with the health care system and laboratory tests.27,28
Although long-term benefits and cost reductions are likely to be greatest for precision medicine efforts directed at the prevention of chronic diseases, the focus of developers and payers so far has been on those tests or therapies that provide short payback periods.29
Finally, there is a basis for concern that the current emphasis on biologic omics will not only divert research energy and funding away from improving traditional approaches to personalized medicine for investigating disease and making clinical decisions, 30 but it will also divert attention away from population health efforts targeting those social determinants of health (poverty, isolation, pollution) and lifestyle factors (smoking, obesity) that drive premature death and morbidity.23,31
Conclusion
Most of the success in reducing morbidity and death over the past half century has been due to earlier detection of disease, public health measures such as smoking reduction and wider application of proven efficacious therapies for primary and secondary prevention. However, the impressive advances in the biologic omics platforms over the past two decades have raised hope that precision medicine will lead to further reductions in morbidity and death. Although it has improved outcomes for selected conditions and the insights emanating from precision medicine efforts hold substantial promise for better disease treatment or prevention,2,32 we are still in the early days of omics-based precision medicine and much has to be done to fulfill that promise.33 The current focus on the biologic omics in discussions of precision medicine should not divert attention from traditional approaches to personalized care: the clinical evaluation, the importance of clinician–patient rapport, and the need to address social determinants of health and lifestyle behaviours. To achieve further improvements in health care, progress on all of those fronts needs to continue, not just in omics-based medicine. Moving forward, we believe that all innovations (including, but not limited to, precision diagnostic, prognostic or therapeutic strategies) should be properly evaluated in robust outcome-based studies. Only those innovations that improve the effectiveness, safety and efficiency of care should be embraced by clinicians and planners in health systems.
Acknowledgements
The authors thank Dr. David Naylor (University of Toronto) for his helpful comments on earlier drafts of this article and the anonymous peer reviewers for their input.
Footnotes
Competing interests: None declared.
This article has been peer reviewed.
Contributors: Finlay McAlister conceived the article and wrote the first draft of the article. Andreas Laupacis and Paul Armstrong critically revised it for important intellectual content. All of the authors contributed substantially to the work, gave final approval of the version to be published and agreed to be accountable for all aspects of the work in ensuring that questions relating to the accuracy or integrity of any part of the work are answered.
Funding: Finlay McAlister is supported by Alberta Innovates – Health Solutions and holds the Capital Health Chair in Cardiac Health Outcomes at the University of Alberta. Andreas Laupacis holds a Canada Research Chair in Health Policy and Citizen Engagement.