Technology: Fluorescence in situ hybridization (FISH)
Use: FISII allows direct visualization of individual chromosomes or genes in tissue and cell specimens and is used as a cancer diagnostic tool to identify large DNA alterations.
History: To a cytogeneticist or anatomic pathologist, "going FISHin'" has quite a different meaning than the usual relaxing pastime. FISII was developed in 1986 as an offshoot of in situ hybridization.[1, 2] In the original technique, a nucleic acid probe (single strand of DNA or RNA) was labelled with a radioisotope and allowed to bind to its complementary strand in a tissue or cell specimen. The location of the signal could be observed by autoradiography on high-resolution photographic film. FISH was developed to identify, by fluorescence, cells containing a genetic target from a complex mixture. The probe is labelled with a fluorescent dye that hybridizes to a DNA target and can be directly seen with fluorescent microscopy. Major advantages of FISII are the brightness of the signal seen and the narrow wavelength of light that each fluor emits, allowing multiple fluors to be easily distinguished in the same sample. Furthermore, the fluor is confined to the binding site, unlike other dyes which precipitate around the binding site. This precise localization allows mapping of the fluorescent signal to subcellular levels and can even be used to determine the location of specific genes (Figure). This technique can also allow quantitation of the number of targets (or genes) in a specific interphase cell nucleus (Figure inset) and may eventually be useful as an indicator of biologic potential in a tumour such as a breast carcinoma. The simultaneous use of numerous fluorescent probes, each emitting light at a different wavelength, makes FISH one of the most aesthetically pleasing of all medical diagnostic tests.[3, 4]
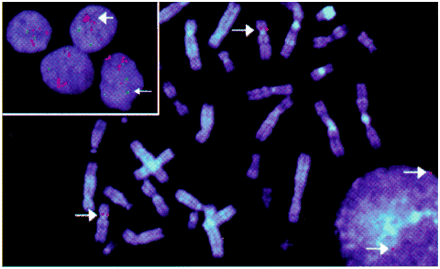
Figure. Visualization of human genes using FISH. A DNA probe for single-copy gene is labelled with fluorescent dye and allowed to bind to normal metaphase chromosome spread (stained blue-white). Interphase nucleus also shows 2 copies of target gene. Inset: Amplification of genes in breast cancer cells. Green signals indicate normal number of copies of RB1 gene, and red signals indicate excessive number of copies of CCND1 (Cyclin D1) gene.
Promise: With its ability to identify single genes FISH can be used in uncultured interphase cells, providing a much faster and cheaper method of chromosome identification than band karyotyping, which depends on living cell cultures. Current clinical uses of FISH involve rapid chromosome counting (e.g., to diagnose Down's syndrome) in interphase cells and identification of chromosomal rearrangements such as translocations in cancer specimens.
Problems: FISH requires significant technical expertise to obtain consistent results and a good fluorescent microscope; a digital camera is highly recommended.
Prospects: More recent applications of FISH include comparative genomic hybridization3 and spectral karyotype analysis.4 In these applications FISH may be more sensitive than conventional karyotyping for detecting chromosomal abnormalities in cancer specimens. Enhancements to FISH occur frequently.[5, 6] Future diagnostic applications may involve quantitation of genes such as c-erbB-2 as predictive or prognostic indicators in cancer specimens. Furthermore, digital image capture and quantitation is ideal for fluorescent images and has the potential to be more sensitive than other visualization techniques. Such imaging technology is also amenable to automated analysis, thus offering the prospect of lower costs, shorter turnaround times and greater productivity than manual methods.
New technologies are emerging such as laser-capture microdissection, with which specific cells can be dissected from a tumour specimen, and array chips, in which hundreds or thousands of molecular parameters may be analysed from a single specimen. These technologies may eventually offer quantitation of a wider panel of specific genes from clinical samples and will likely decrease some of the diagnostic applications of FISH.
Competing interests: None declared.