Our first commitment as clinicians is to our patients as individuals. This understanding lies at the heart of the precision medicine initiative, which aims to customize treatments based not only on a patient’s clinical picture but also on their genetic, demographic and environmental profile.1 Clinical trials are rigid by design, typically assessing the effect of one intervention. In a reality of complex patients and polypharmacy, clinical trials cannot provide all the answers that clinicians are seeking. It is widely accepted that the uniquely complex health care environment requires the implementation at scale of informatics tools to help physicians navigate through the huge volume of medical data, and technology is being developed and refined to support this need. Whereas data from electronic health records are routinely reused in research to generate evidence-based recommendations for patient care, databases of clinical trial populations are not. The potential of individual-level data of participants in completed clinical trials to facilitate evidence-personalized treatment decisions has not yet been fully realized.
Problems with current models of evidence synthesis
Facing a rapidly growing body of medical knowledge, physicians rely on high-level aggregate-based knowledge resources, such as online knowledge bases, meta-analyses and clinical practice guidelines. However, this results, by default, in clinicians treating individual patients as if they were members of a homogeneous group. Yet, any successful trial is likely to have some participants who do not do as well as the “average” participant; what was best for the group may actually have been harmful for those participants. The masking of between-participant variability by aggregate-level design and analysis complicates prediction of the utility and risk of applying a given intervention to a particular patient.2
A meta-analysis may conclude that an intervention’s net (mean) effect is negligible, which would justify avoiding that intervention. However, if considerable variability exists among the populations and results of the analyzed trials, this conclusion may not apply to an individual patient who differs from the average patient. Frequent use of restrictive eligibility criteria in trials may further compromise their relevance to a particular patient, because they result in trial populations that do not properly represent that patient. This may limit the external validity of those trials3 with regard to a target population or a specific patient.
Providing all patients with the best-on-average treatment is arguably more justifiable than not relying on evidence at all, but the interests of the individual patient may not be well-served. Precision medicine attempts to close the evidence gap between a generic and real patient.4 Indeed, differentially applying evidence to different patient populations is an established practice, notably in cancer and coronary heart disease therapy. Subgroup analysis of clinical trial populations is commonly used to support this strategy. However as subpopulations are typically characterized by categories of a single baseline characteristic (e.g., age range or sex), a patient may belong to multiple subgroups exhibiting discordant trends with regard to the effect of an intervention. Integrating subgroup analyses to support clinical decisions related to a particular patient is a challenging task.5 Although there may be extensive evidence on the optimal management of a particular condition, using this management plan for patients who have multiple comorbid conditions may not be well supported. Dörenkamp and colleagues term this “the maze of multimorbidity.”6
Physicians commonly use clinical judgment when adjusting evidence to the care of an individual patient based on the patient’s special characteristics and circumstances. Unfortunately, the evidence gap commonly reduces this to a subjective (and thus often biased) process.7
Alternative approaches to using accumulated data to inform care for the individual patient
Matching patients to the right trial
Preferential reliance on clinical trials with study populations similar to the patient at point of care is one way that evidence can be applied to the individual (Appendix 1, available at www.cmaj.ca/lookup/suppl/doi:10.1503/cmaj.160267/-/DC1). Some existing tools and resources could be adapted to facilitate picking the “right” trial(s) by matching patients with already completed clinical trials. Open clinical trials are listed by the US National Institutes of Health (available at ClinicalTrials.gov) and its European Union counterpart, the EU Clinical Trials Register (available at https://www.clinicaltrialsregister.eu/ctr-search/search), and tools have evolved to use these databases to match patient profiles with the eligibility criteria in the trials. The American Cancer Society offers a free Clinical Trial Matching Service for patients with cancer.8 Other commercial tools are available (e.g., Emerging Med9 and IBM Watson for Clinical Trial Matching10), and these could be adapted to the purposes of precision medicine.
Using individual-level trial participant data
If the individual-level data of participants in clinical trials were available, they could be used to individualize the results of clinical trials. For a patient of interest, a subpopulation of trial participants with similar baseline characteristics could be formed. The outcomes of that subpopulation could then be analyzed to estimate a personalized risk–benefit ratio for an intervention studied in the trial.
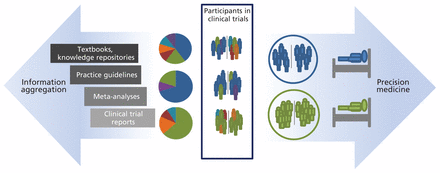
Moreover, similar participants from across different but comparable clinical trials could be pooled to form an ad hoc retrolective cohort11 (Figure 1). Using this approach, analysis of data of similar individual participants of trials could inform a personalized choice of drugs and dosing regimens.
Traditional evidence-based medicine is practised based on increasing levels of information aggregation (left arrow). Similarity matching at the individual trial-participant level supports the formation of retrolective cohorts to inform precision medicine (right arrow).
How could this be operationalized? Similarity search is already used in a variety of domains, from plagiarism detection to cluster analysis. Similarity assessment of nucleic acid sequences and proteins (e.g., used by the Basic Local Alignment Search Tool [BLAST], available at the National Center for Biotechnology Information)12 has proven very useful in biomedical research. In the clinical setting, the use of similarity-driven computer-aided diagnosis systems in fields such as radiology is growing.13
Genomic data, microbiota signatures and proteomic profiles are starting to be included in the electronic health records of patients. Although such data can greatly improve our understanding of patients, they are also overwhelmingly detailed and difficult to interpret even for expert clinicians. However, those data could allow for more clinically meaningful similarity analyses to be computed. A cohort of trial participants similar to a patient in question that is based on more clinically meaningful metrics is likely to result in more reliable outcome predictions.
In some cases, recommendations based on such analyses of individual patient data are likely to deviate from the general guidelines.
What are the challenges in similarity assessments?
Reuse of individual patient data from clinical trials in advanced similarity assessments is challenging both conceptually and practically. Combining multiple data types to assess similarity at the patient level as a whole is complicated. Only baseline characteristics of trial participants that are collected can be considered in the matching process, and those are typically a small subset of the parameters that could be used to characterize an individual. However, trials do collect a wide range of demographic (e.g., age, sex and race) and clinical (e.g., disease severity and comorbidities) features, as well as setting-related features (e.g., type of hospital and geographic location).
Characteristics that predict the intervention outcome should be included in the similarity assessment. Unfortunately, these remain largely unknown and may vary by disease, intervention and subject. Indeed, domain knowledge can help in selecting relevant characteristics. Based on a physician’s clinical knowledge and judgment, they may define, on a case-by-case basis, through an interactive computer program what features should be used to select similar patients. Although of questionable validity, this is what physicians currently (and often implicitly) do when assessing the overall similarity between a patient and a trial population. Appreciating that physicians’ limited knowledge may not be sufficient to form a valid and comprehensive list of relevant characteristics, an alternative approach would be to include all available characteristics in the similarity assessment.
Some features that are relevant to similarity assessment may be more important than others. In most cases, it would probably be impossible to assign differential, evidence-based weights to various features. As with feature selection, clinical judgment could be used to weigh feature importance or unweighted assessment could be applied.
The use of individual-level data, especially with new data types, raises privacy issues, because integration of multiple data types has the potential to be used to identify patients. One way to ensure privacy is to mask individual-level data by only allowing users to view aggregated characteristics of a group of similar patients. For example, in the Observatonal Health Data Sciences and Informatics program, 14 clinical queries are run on disparate participating clinical databases and return aggregate-level results. This enables researchers across institutions and states to leverage insights from the data of millions of patients without exposing patient identity.
In some cases, sufficient data are not available to guide the management of a patient with a rare disease or unusual circumstances. Similarity analysis could be used to collect such cases, and the data would be used by clinicians in the same manner as case reports.
Evaluation of information resulting from automated analysis of clinical trial data would require human judgment. Some factors are important when making clinical decisions that are very difficult to measure and report in clinical trials. Patient frailty, values and preferences; availability of support at home; physician experience and beliefs; and other considerations influence the translation of evidence to real-world decisions.
How can statistical inference be used to personalize clinical trials?
Careful statistical analytic practices are imperative to support inference from subgroup analysis.5,15 A subgroup of similar patients will be smaller than the original population from which it is selected. Inevitably, the precision of estimates of intervention effect size will be affected. This is the case with both demographic and clinical characteristics including comorbidities.16 Certain populations, especially older adults, infants and children, sicker patients or those with comorbidities, are generally underrepresented in clinical trials.17,18 Subgroups of trial participants having those characteristics are likely to be smaller and thus less informative. Multiple post-hoc analyses are associated with a risk of false-positive findings, requiring the use of smaller p-value cut-offs for statistical significance.18,19 Considering that subgroups are smaller and thus have less power, reaching statistical significance when participants are drawn from a single clinical trial may be difficult. Nevertheless, under some circumstances (e.g., when a subgroup differs substantially from the general trial population), such analyses may produce more accurate estimates, which is why subgroup analyses are done. As with meta-analyses, pooling similar participants from across multiple comparable clinical trials may increase the subpopulation sample size and narrow down confidence intervals of the estimates of intervention effect.
Moving forward
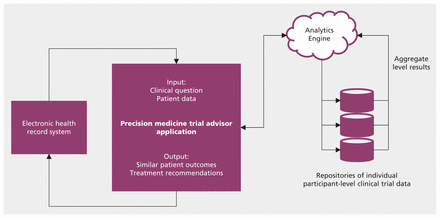
Figure 2 illustrates a high-level architecture of a hypothetical computer application to support precision medicine by personalizing the results of clinical trials. An example of a clinical use of this application is illustrated in Box 1. Some of the infrastructure required for this type of application to operate has already been developed.14 Data from electronic health records provide the input needed to automate the matching of patients and trial participants using computable similarity scores. Methods for extracting data from individual patient records across different systems have been implemented.14 A consortium of pharma companies (ClinicalStudy DataRequest.com) is making clinical trial data at the individual participant level available for researchers. However, these data remain unavailable for clinicians to access at the point of care. Regulatory issues and the financial interests of stakeholders will need to be carefully considered to facilitate access to such data at scale. Methodologies using similarity search that have been developed in other domains will have to be adapted to handle clinical data, and their performance will need to be validated. Similarity-based prediction tools can be heavy consumers of computational power. Investment in processing and communication infrastructure is likely to be required to accommodate their use.
High-level architecture of a framework for providing individualized evidence from clinical trial results. A clinician uses a precision medicine trial advisor application to retrieve evidence for a patient in current care. The application extracts relevant clinical data from electronic health records. A cloud-based analytics engine queries individual participant-level clinical records across respositories of trial data and collects aggregate-level results, which are analyzed and sent back to the application for the clinician to review.
A case for using a hypothetical, computer-aided and similarity-based analytics application for personalizing the results of clinical trials
A physician is evaluating a 60-year-old white man with coronary heart disease and low high-density lipoprotein cholesterol, who does not have diabetes. The patient has high low-density lipoprotein cholesterol but is intolerant to statins; therefore, the physician considers adding a fibrate to his treatment.
Using a Web application interface, the physician types in the keywords “fibrates” and “coronary heart disease” to find relevant clinical trials. The physician reviews a shortlist of article titles and abstracts to select trials that assess fibrates for primary or secondary prevention. The physician selects outcomes of interest and patient baseline characteristics to be used in the analysis. Common data models are used by the application to align metrics across trials.
The system then automatically extracts the selected patient characteristics and runs a similarity analysis against the baseline characteristics of individual trial participants, forming a cohort of participants similar to the patient. The outcome rates of cohort participants are calculated by trial arm, and adverse reactions are ranked. The system performs statistical analyses and presents the results to the physician, including a comparison with the results obtained for the entire population studied.
The physician then evaluates the results and discusses possible treatment options with the patient.
Published information is more reliable for guiding the care of average patients than supporting decisions that relate to patients diverting from the average.
By applying automated similarity-based matching of a particular patient to clinical trial populations and individual participants, insights relevant to that patient may be derived from the course of disease in similar patients.
There are compelling arguments for reusing clinical trial data to offer patients more precise care, but there are also substantial theoretical and practical barriers, and concerns about the utility of this approach.
Footnotes
Competing interests: None declared.
This article has been peer reviewed.
Contributors: Amos Cahan conceived the idea and drafted the manuscript. James Cimino provided important intellectual content, and critically evaluated and revised the manuscript. Both authors approved the final version to be published and agreed to act as guarantors of the work.
Funding: Amos Cahan was supported by an appointment to the Research Participation Program for the Centers for Disease Control and Prevention, National Center for Environmental Health, Division of Laboratory Sciences (DLS), administered by the Oak Ridge Institute for Science and Education through an agreement between the US Department of Energy and DLS. Amos Cahan is currently employed by IBM. James Cimino was supported in part by research funds from the National Library of Medicine and the National Institues of Health (NIH) Clinical Center, and is currently supported by the Informatics Institute at the University of Alabama School of Medicine, Birmingham.